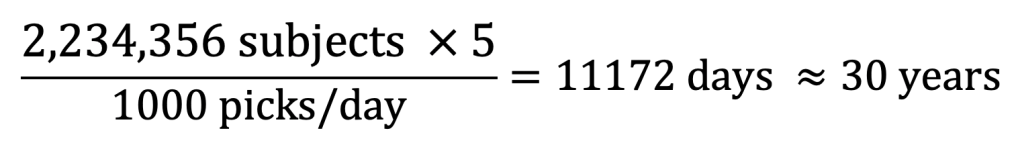Mito Mapper workflow (Top View) is complete!
As you might have noticed, we have entered the second stage of this project – the Mito Mop Up workflow, in which you are invited to clean up the locations picked during the Mito Mapper workflow.
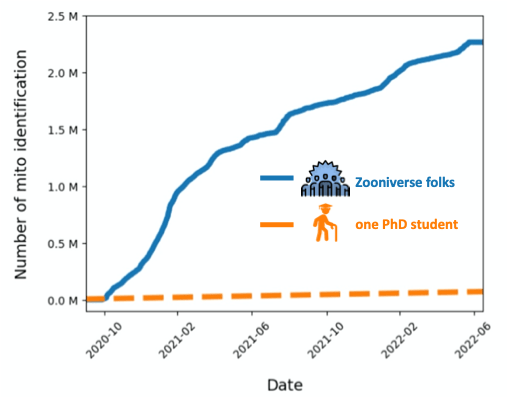
First and foremost, we would like to thank all of you for your amazing work so far. In the Mito Mapper workflow, you have created a whopping 2,234,356 identifications! The plot below shows how we got here. If a PhD student identifies 100 mitochondria in a day, it would have taken them 54 years to reach the same number.
Here is a slice from the 3D image stack of placenta acquired by electron microscope overlaid with a distribution of classifications. You did not miss any region that could have a mitochondrion! This large dataset you helped create will be used in the future to develop an automated pipeline for the segmentation of the placenta based on machine learning. The more training data that is available, the more accurate the predictions on new, unseen data will be.
Why are we doing the Mito Mop Up workflow?
At the same time, the accuracy and effectiveness of a machine learning model depends heavily on the quality of the training data. If the training data is incorrect or biased, the model will learn the wrong patterns or relationships from the data, which can result in erroneous or misleading predictions.
For our data, as you probably have noticed, mitochondria are difficult to identify. With their varying shapes and sizes, they can be elusive sometimes even to the eyes of the professionals. This is exactly why we are using the Mito Mop Up workflow. We zoom in and circle every classification made in the Mito Mapper workflow for you to double check whether it is a mitochondrion or not. We have set the retirement limit of this workflow to be 5, meaning that each zoomed in classification will be shown to five volunteers, and the outcome is dependent on the collective choice of the majority (i.e., 3 or more).
Why does the Mito Mop Up workflow need changing? And How?
That sounds like a plan, right? However, we simply could not have predicted that over 2 million classifications would be made in Mito Mapper. We are getting a lot of help for the Mito Mop Up workflow right now, averaging 1000 picks in a day. But then we did the maths and found that it would take 30 years for all the subjects to be checked by 5 volunteers! That’s taking a little too much of everybody’s time.
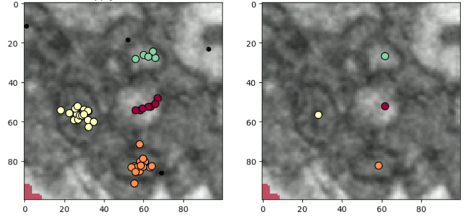
After examining the distribution of the classifications from Mito Mapper, we came up with a solution. In this small region below, there are a few mitochondria-like structures. Most of the picks are on or near them, with a few lone points elsewhere. If we zoom in on every pick and upload it to the Mito Mop Up workflow, the same structure would be shown to tens of volunteers, and we don’t think that is the best use of your valuable time!

Since these picks are spatially adjacent to each other, we can run an algorithm to group them together. We employed DBSCAN (density-based spatial clustering of applications with noise), an algorithm that works by defining a radius around each point and then counting other points within that radius. Based on the number of neighbours within the radius, points are considered to be either a part of a cluster or an outlier. Using DBSCAN, we can group these picks into four nicely defined clusters. If we only zoom in on the centres of these clusters in the Mito Mop Up workflow instead of each pick, it can greatly speed up the process without losing useful information.
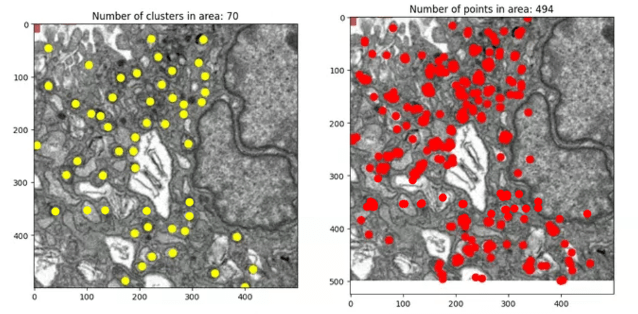
Here’s how the algorithm works on a larger region. The image on the right shows where the picks from Mito Mapper are, and on the left shows the centre of clusters after performing DBSCAN, the number of which has reduced seven-fold compared to the former:
What’s next?
By applying spatial clustering, we can reduce the timeframe of completion for the Mito Mop Up workflow from 30 years down to something far more reasonable. However, your contribution to the Mito Mapper workflow provided even more information. For instance, in the small region we showed above, there are already so many picks on this region that checking up on it again seems unnecessary.
Based on the clustering result, we introduced cut-offs based on cluster sizes to the divide results from Mito Mapper into three categories:
- There are many picks on or closely around this structure. It’s mostly likely a mitochondrion, and people will probably say so if they see it in the Mito Mop Up workflow.
- There are a few picks on this structure. It’s hard to tell whether it is a mitochondrion or not.
- There is only one pick on this structure, even though it has been shown to at least 16 [PS1] volunteers already. It’s unlikely to be a mitochondrion, and people will probably say no if they see it in the Mito Mop Up workflow.
For number 1 and number 3 we can be fairly confident that we know what everyone will answer if they see this structure in the Mop Up workflow. Therefore, we most likely don’t need to double check. However, number 2 is more tricky. Not only this, but we also have the data from the current Mop Up workflow to factor in: have people already seen the structure in 2? And how many of those people have already said yes or no? By using the clustering results and the >480,000 classifications already in the Mop Up workflow together, we hope we can optimise the workflow so that we don’t miss anything out, but we also don’t include things that we are already confident in.
We are working on optimising the Mito Mop Up workflow. However, your current classifications in this workflow will definitely not go to waste! They not only provide valuable scientific information, but will also be analysed and utilised in the optimisation process.
Thank you once again for your time and dedication, and stay tuned for our updates in the future!